Rewriting The Genomic Alphabet
25:58 minutes
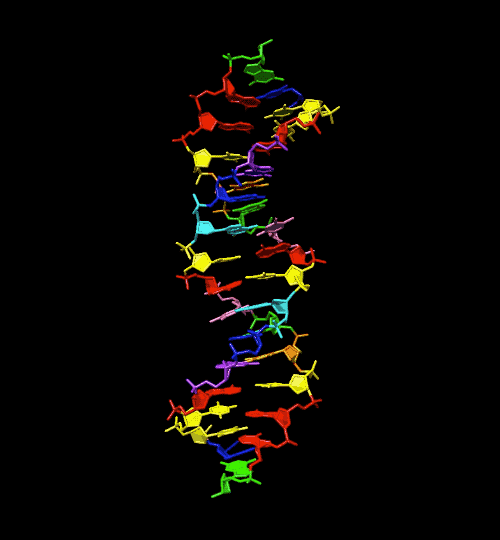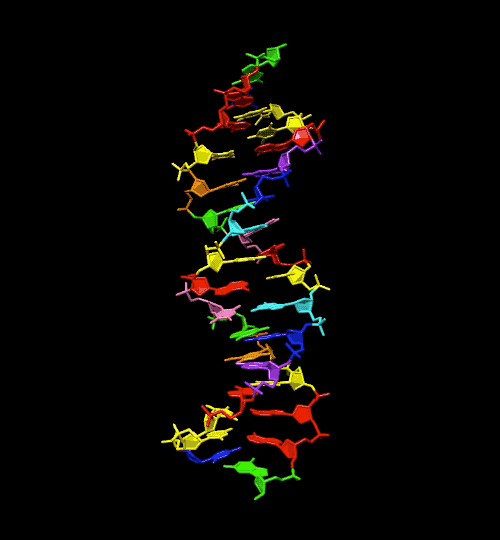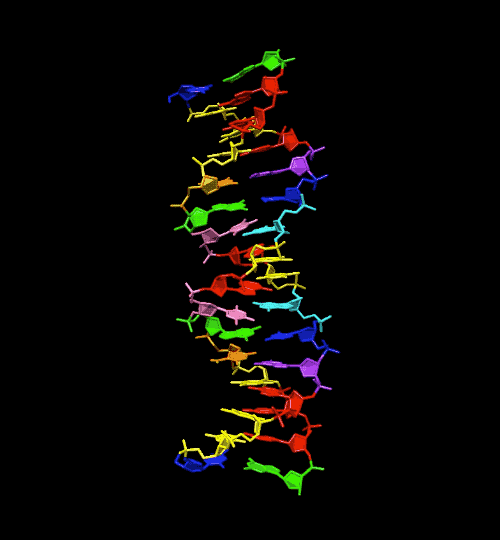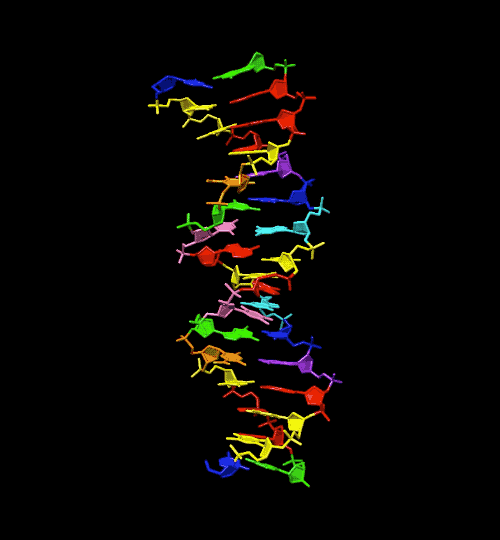
DNA is the universal programming language for life, and the specific code to that program are the combination of the base pairs adenine, guanine, cytosine and thymine. But are those the only base pairs that could be used to create DNA? Scientists looking into this question were able to create 4 different base pairs that don’t exist in nature. Their results were published in the journal Science.
Chemist Floyd Romesberg, who has created his own unnatural base pairs, biologist Jef Boeke, who is working to create a synthetic yeast genome, and bioethicist Debra Mathews talk about how altered genomes could be used for creating novel medicines and fuels—and whether this is considered a new form of life. (None of these guests were involved in the Science study.)
Floyd Romesberg is a professor of Chemistry at the The Scripps Research Institute in La Jolla, California.
Jef Boeke is director of the Institute Of System Genetics and a professor in the Department of Biochemistry and Molecular Pharmacology at New York University Langone Health, in New York, New York.
Debra JH Mathews is assistant director for Science Programs at the Berman Institute of Bioethics, and an associate professor of Pediatrics at Johns Hopkins University in Baltimore, Maryland.
IRA FLATOW: DNA– well, you’ve heard this your whole life– it is the programming language of our cells. And that the four base pairs, known as A, G, C, T, make up the code of DNA. But could our DNA be built on a different code, maybe more letters in the code. Researchers have built unnatural base pairs now. And they’ve added them to the genetic alphabet, doubling the number of base pairs to eight in some cases– in some of the research. The results were recently published in the Journal of Science.
This is one way to retool DNA. Other scientists have been trying different types of base pairs, trying to build genomes from scratch. All kinds of new products are possible– medicines, fuels. What could go wrong, right? Haven’t we heard this before? That’s what we’ll be talking about next. The brave new world, literally, of synthetic genes– the good, the bad, and the who knows.
Let’s open the discussion about technology with my next guests, both deeply involved. Dr. Floyd Romesberg is Professor of Chemistry at the Scripps Research Institute in La Jolla, California. Hi, Dr. Romesberg.
FLOYD ROMESBERG: Hi, Ira.
IRA FLATOW: Dr. Jef Boeke is Director of the Institute of Systemic– I’ll get it right, Jef– System Genetics, and a professor in the Department of Biochemistry and Molecular Pharmacology at New York University, Langone Health, here. Welcome back, Jef. Good to see you again.
JEF BOEKE: Good to see you.
IRA FLATOW: Well, just to define terms– how would you define synthetic genome to someone who has never heard of it before?
JEF BOEKE: Well, a synthetic genome– the way we look at it is a genome that’s built starting from non-living matter, and can be designed on a computer, and introduced into a living cell to essentially reprogram that, like upgrading your operating system from the old to the new. You can add new features to the genome by changing the DNA sequences in specific ways. But the recent research that you highlighted goes way beyond that in actually adding new letters to the alphabet that we have to play with.
IRA FLATOW: Is that something a lot of people are aiming to do?
JEF BOEKE: Well, I think it’s a small and very select group of people that are carrying out that type of research, and it has all kinds of interesting challenges. But it is very exciting to see that expansion of the alphabet, at least in a test tube.
IRA FLATOW: Floyd, you weren’t part of this latest study, but you have created your own unnatural base pairs in your lab, right?
FLOYD ROMESBERG: Yeah, that’s right.
IRA FLATOW: Tell us why you do that and why this latest study is such a big deal?
FLOYD ROMESBERG: Sure. So why someone would want to do that? So the natural letters, as you already mentioned, G, C, A, and T, they’re what cells use to make proteins. Proteins are what cells use to do what they do. So everything that you do is largely carried out by the proteins that you’re able to make.
But it turns out that those natural four letters encode for generally only 20 amino acids, sort of canonical natural amino acids. And those amino acids don’t actually do some things that we might want them to do. They’re actually– if you went to a medicinal chemist and said, is there anything interesting here, they’d be less than inspired compared to the sorts of molecules they build where they include different functionalities that help make drugs, for example, be the drugs they are.
So what we tried to do, what we set out to do 20 years ago, was to create a new base pair. Instead of four letters and two base pairs, we wanted to make six letters and three base pairs. That would increase the information that you could store, give you new information in a cell that the cell could retrieve and make new proteins with, and maybe proteins designed to have properties like, for example, being drugs to treat diseases that were formerly untreatable.
So this latest work by Steve Benner, it’s a little bit different. Steve’s a good friend of mine. I have a lot of respect for him. In fact, he’s been doing this longer than I have. And his work was a real inspiration to me when I was starting my own lab. But what Steve has done is he’s doubled these letters, but it’s only in a test tube. And what we tried to do was something a little bit different. What our goals are, are to have these extra letters where they can be replicated by DNA polymerases. And not only in a test tube, but in a living cell where they can be used to produce proteins.
What Steve has recently done in this paper that you mentioned was create these four additional letters that he’s been optimizing for quite a long time, and simply looked at their stability in terms of forming duplex DNA. So they’re recognition single strand of DNA forms a duplex that pairs together, and he’s shown that you can make those same pairings with these four extra letters.
IRA FLATOW: I’m Ira Flatow. This is Science Friday from WNYC Studios. Talking with Dr. Floyd Romesberg and Dr. Jef Boeke. Have they been functional? Do these extra DNA– if you’re using these six pairs, can they reproduce and make themselves over again?
FLOYD ROMESBERG: So it’s six letters and three pairs. And so in our case, in my lab, we now have what we call semi-synthetic organisms. They’re just bacteria right now– E. coli. They live and they grow, and they divide, and they maintain those extra letters in their DNA. They can transcribe them into what we call mRNA and tRNA. Those are the intermediate functional versions of DNA that go out into the cell and make proteins. And we’ve, in fact, made proteins with unnatural amino acids. And I started a biotech company called Synthorx, and they’re using the technology to develop some anti-cancer drugs, that is really exciting to watch.
IRA FLATOW: Jef, do you think an entire organism could be built this way with unnatural base pairs?
JEF BOEKE: I think that we’re really a long ways off from that for more than one reason. The current three base pair system that Floyd talked about– it’s amazing what his group has been able to do, but as far as I understand it’s only possible to put one of those special base pairs in a row, whereas the natural genetic code allows you to put any number of them in a row. So there are limitations to the systems that exist right now that might make it hard to really program an entire organism. Even more daunting is the fact that we barely know how to program an organism from scratch with the four natural bases.
So I really think that that’s still very far on the horizon, not to mention the fact that the eight base system that was recently described is only working at a test tube level right now, and not as Floyd said, inside living cells. So call me the conservative in the group. The four base pairs of DNA are plenty wild and crazy for us to work with right now. But we’re not creating truly new organisms for this. We’re sort of endowing existing organisms with new functions.
IRA FLATOW: Our number 844-724-8255, if you want to call us. You can also tweet us @scifri. But even using and– can you rearrange things, the base pairs? Your eyebrows went up when I said that, Jef.
JEF BOEKE: Well, yes, we’ve taken that to an extreme recently. So we had a paper last year where we had a lot of fun with the yeast genome. Now, the yeast, which is our favorite organism of choice, has its genome packaged into 16 units called chromosomes. And I’d like to just mention an analogy I’ve been thinking of for what we did. So if you think of– I don’t know how many people remember what encyclopedias are– but an encyclopedia is a series of volumes that has a whole bunch of information in it. And it’s a good analogy for a genome. The volumes would be the chromosomes.
IRA FLATOW: I’m going to stop you there because we have to take a break. And we’ll stop at the encyclopedia, think of your books, think of them how they’re lined up. We’ll be back to talk with Jef Boeke and Floyd Romesberg. And Debra Mathews is going to join us to talk a little bit about the ethics of all of this work. So stay with us. Our phone number 844-724-8255. We’ll be right back after this break. Don’t go away.
This is Science Friday. I’m Ira Flatow. We’re talking about synthetic genomics with Floyd Romesberg and Jef Boeke. And Jef was talking about building a completely synthetic genome in yeast. You got us up to an encyclopedia with a lot of books in it.
JEF BOEKE: Right. So the genome can be thought of as an encyclopedia consisting of 16 volumes. And essentially, what we realized was we could rip off the back cover of the first volume of the encyclopedia and the front cover of the second volume, and then glue those two volumes together. And by doing that over and over again, we created, essentially, a genome on one giant chromosome.
And it has all the same information in it, so it’s still useful. It’s a little bit more tedious to handle. But remarkably, the cells took this very much in stride and didn’t complain very much. And that was a very surprising result to a lot of people that you can really rearrange the DNA sequences in such a dramatic way and still have everything work more or less normally.
IRA FLATOW: Let me bring on someone else who can talk about and react to the research that’s going on here. I want to bring on Dr. Debra Mathews, Assistant Director for Science Programs of the Berman Institute of Bioethics, and Associate Professor of Pediatrics at Johns Hopkins University. Welcome to Science Friday.
DEBRA MATHEWS: Happy to be here.
IRA FLATOW: You’ve been listening, right?
DEBRA MATHEWS: Yep.
IRA FLATOW: What’s your take on all of this?
DEBRA MATHEWS: Which piece? I mean, first, I want to emphasize something that Jef said earlier, which is that this is all at a very early stage. Whether you’re talking about the hatch emoji DNA, these four new bases– with the best name ever, by the way– or whether you’re talking about Jef’s synthetic genomes, they’re fairly basic. But we need to think about, I think, what we do going forward. What these technologies mean for us now for science? What they can do for us? Lots of great potential applications, like new drugs, new vaccines. Lots of really interesting basic science. But potentially, risks as well.
Floyd mentioned that he’s working in E. coli. And E. coli was the organism that kicked off our worries about recombinant DNA. So when we first, in the 1970s, were able to cut and paste DNA. The experiment that was done that freaked people out, scientists and the public alike, was introducing an antibiotic resistance gene into E. coli. E. coli lives in all of our guts. And that worried people. We didn’t know then what the risks would be. And so the scientific community asked for regulation, both because they were concerned about safety, both for themselves and the general public. And also, because they understood that this was controversial.
IRA FLATOW: Well, let me ask my scientists here. Jef or Floyd, how do you control the synthetic genomes to make sure that the organisms when you put these genes all together, and they’ve never been together before, make what you think they’re going to make?
JEF BOEKE: Right. Well, we take a number of precautions. We tend to work on the yeast front– of course, it is naturally a very safe organism– but we work with strains that carry mutations, that essentially render them unable to compete well in the wild. And we can introduce much more extreme forms of control. It’s actually an area we’ve worked on because we take this matter of safety of engineered organisms very seriously. And I know– I mean, in Floyd’s case, I think he’ll have a lot more to say about that. With an extra base pair, you can exert a special kind of control. So maybe I’ll hand it over to him.
FLOYD ROMESBERG: Yeah, that’s right. If I could just inject– so our X and Y are fifth and sixth letter that form that third unnatural base pair. Like Debra just said, we have it working in E. coli. We’re working on getting it working in other organisms and other cell types. But we have to provide X and Y– the precursors of X and Y– to the cell. And this isn’t a Jurassic Park-like situation, because life’s not going to find a way. These are increased complex biological molecules that nature doesn’t make. They don’t make anything like them.
They each would take something like 15 new steps based on enzymes that nature doesn’t have anything like right now. And the notion that nature could out of nowhere assemble such complex pathways it’s just a dauntingly– it’s just a very unlikely thing. I’m myself– I’m going to catch all sorts of flak for this– I’m comfortable calling it zero. So we have in our case a built-in safety system, because if we don’t provide the precursors of X and Y to the cells, they can’t maintain them in their DNA. And so if they escape into the wild, they’ll die. And we’ve shown that many times.
IRA FLATOW: Debra, are you satisfied?
DEBRA MATHEWS: No, I mean, as I said, this is very basic right now. And we’re actually in a even safer place, because there’s very strict control over who has access to the new bases. It’s not like you can just buy them. Or can you?
[LAUGHTER]
FLOYD ROMESBERG: You actually can.
DEBRA MATHEWS: You can. Great.
FLOYD ROMESBERG: Some of them you can.
DEBRA MATHEWS: Some of them you can. So there’s more control over the technology now, I would imagine than there will be in the future. And I think we’re in a really strong position to think about what this does look like going forward.
IRA FLATOW: What do you mean you can buy them?
FLOYD ROMESBERG: Well, so actually, this wasn’t even my lab. A few years ago, I was sitting at my desk and I got a phone call from a company that makes nucleotides. That’s our name for these letters. That’s what scientist call them. And they actually said, well, we have a question about a paper you published and the synthesis of one of these analogs. Could we ask you a question? So I obviously knew nothing about it, because I sit at a desk and I’m fortunate enough to work with some really talented people who actually do do things. So I had to go get my co-worker and then we explained it to them.
And in this process we found out that they had based on our papers decided to make them and sell them. So there’s actually a couple of different companies now that sell some of the analogs. And they could make a functional semi-synthetic organism. But in their hands, it also will not be able to escape.
IRA FLATOW: But they can do their own work on it?
FLOYD ROMESBERG: Yeah, and several labs are.
DEBRA MATHEWS: Mm-hmm.
IRA FLATOW: Jef, can we buy your yeast?
JEF BOEKE: You can’t buy it. But if you work at an academic institution, you can certainly obtain it. We don’t really charge money for it.
IRA FLATOW: I don’t know.
[LAUGHTER]
I feel a little queasy about that. Debra, do you share my queasiness?
DEBRA MATHEWS: Yeah. I certainly would want there ideally to be a process in place to vet who has access, to think about what the possible uses of these new nucleotides might be in the organisms that take them up. The fact is we don’t know how our system would react to novel nucleotides necessarily. And an issue that almost always comes up when you’re talking about synthetic biology generally is that of dual use.
These scientists are doing fantastic, creative work with the best goals in mind, but not everyone in the world has the best goals in mind. So we might design something for very good purposes, but someone else might use it for bad purposes. And if, for example, these new nucleotides were able to make drugs more potent or more persistent, which would be great in terms of treating disease. But more potent and more persistent would be bad in the context of a really harmful disease.
IRA FLATOW: Let me go to the phones. 844-724-8255. Let’s go to Patrick in Yardley, Pennsylvania. Hi, Patrick.
AUDIENCE: Hi, how are you? Yeah, thank you. Great subject. My question is, how does this research intertwine with CRISPR technologies? And how will it enrich CRISPR in the future?
IRA FLATOW: Jef, what’s the difference between this and CRISPR? It’s a good question.
JEF BOEKE: So CRISPR is an extremely useful tool that most of us use. I know Floyd has used it in interesting ways as well. So a lot of the work we do in yeast actually can be done without CRISPR, and I would say 95% of the work we do with our synthetic yeast genome is done without using CRISPR. But CRISPR is an amazing tool. It can be deployed in yeast, as it can in many other organisms.
IRA FLATOW: If and when you fully synthesize the yeast, is it another form of life?
JEF BOEKE: Well, we call it SE 2.0, so it has a different name. But it can interbreed with Saccharomyces cerevisiae the natural counterpart. So it is formally part of the same species. So I would say no, by the biological species definition, it’s not a new species.
IRA FLATOW: But would it be a new living thing?
JEF BOEKE: Well, remember, it’s just a form of this brewer’s yeast. And there are many forms of it out in nature. This one has engineered components in it and has a lot of engineered components in it. So it is able to do things like rearrange its genome on command that the natural yeast cannot do. But again, by the traditional species definitions, it’s part of the same species.
IRA FLATOW: So I take that as a yes then.
JEF BOEKE: That depends what the question was.
[LAUGHTER]
IRA FLATOW: Is it alive? Is it a form of life?
JEF BOEKE: It’s a new form of life, absolutely. But it is part of the species we know and love as brewer’s yeast.
IRA FLATOW: Floyd, can you take out everything from a cell and put all your synthetic genome in there? And now we have a cell with something totally different for the nucleus.
FLOYD ROMESBERG: So as Jef just alluded to a few minutes ago, no, you can’t do that. We’ve taken very much, in some respects, the opposite approach, where we’re starting in a top-down approach. Where we’re going into an organism that has a full genome. Our base pairs– our fifth and sixth letters that pair to form a third base pair– they pair in a completely different way than nature’s base pairs do. And they actually do exert a small distortion on the duplex. And they do rely on being flanked by natural letters.
Now, we’re actually just publishing some work that shows that we can actually put several in a row and we can actually put a pretty high density, such that you wind up getting amino acids in a protein that are right in a row. But, for example, they’d be separated. We can put two in a row in DNA and still have cells replicate it and hold onto it. We can put many as long as they’re separated by one or two.
But the way this works is these letters form codons. These codons are what determines proteins. These codons are combinatorial. So in order to encode a new protein, you don’t need them to be replicated in a row. Nature has existed– life has existed to this point by synthesizing every one of the proteins it’s ever made with 61 codons. That’s these words that can– and we now have 64. We’ve now added three.
We’re about to publish a paper where we now have our E. coli cells decoding three new codons. And to do that kind of thing– and there are just so many fascinating applications that one could think about applying this to– it is certainly correct that we can’t put long strings in a row. But we also don’t have any applications that would need that. And there are just so many neat things that we can do the way that we’re doing it.
IRA FLATOW: I’m Ira Flatow. This is Science Friday from WNYC Studios. Running out of time, unfortunately. Because you were the last one to mention the neatest thing, what would be, Floyd, the neatest thing that you can do? You said you can do so many neat things.
[LAUGHTER]
Quickly.
FLOYD ROMESBERG: I’ll tell you right now the two things that we’re working on. A collaboration with this company, Synthorx, that I mentioned, we’re actually making an aisle two drug that is really pretty impressive. And the chance to watch them possibly go cure incurable diseases will be just so gratifying. And secondly, we’re going to make organisms that instead of providing proteins to us to do things, we’re going to make new organisms that they themselves do new things. For example, like eating oil spills and digesting them down to carbon dioxide, hydrogen in water.
IRA FLATOW: So these would all be patentable new organisms then?
FLOYD ROMESBERG: That is correct.
IRA FLATOW: Floyd, where the rubber meets the road, so to speak?
[LAUGHTER]
FLOYD ROMESBERG: Well, so some of the work done in my lab is funded by this company that I mentioned, and they are, of course, interested in funding it if they can own it.
IRA FLATOW: Jef?
JEF BOEKE: Yeah, I think my last word would be it’s really important for us to think about the things that might go wrong, but we should also really think about the things that could go right and there’s a lot of them.
IRA FLATOW: Debra?
DEBRA MATHEWS: Yeah, there’s certainly potential opportunity costs, but there are also values beyond scientific values that are important. And the public should have an opportunity to think about and talk about and voice their own values about how this technology ought best be put to use.
IRA FLATOW: That’s what we’re doing here today. One last thought before I want to leave you. If you can come up with new kinds of synthetic DNA, does that mean if we go to other planets, we should be ready to find things we did not expect to find if we find living things? Jef?
FLOYD ROMESBERG: Yes.
IRA FLATOW: Floyd?
FLOYD ROMESBERG: That was me. Yes.
IRA FLATOW: And Jef?
JEF BOEKE: I think it’s possible. But Francis Crick actually was a proponent of the idea of panspermia, which is that our DNA bases actually came from outer space. They’re floating around throughout the cosmos and they’re a universal. So I think that’s a formal possibility we should also consider.
IRA FLATOW: Debra, are you ready to find new life?
DEBRA MATHEWS: Absolutely.
IRA FLATOW: Do you think it would radically change the way we look at life here on Earth?
DEBRA MATHEWS: Well, I think it’s incredibly interesting that there are these other bases that can be made and can function in systems. But we ended up with the four we ended up with.
IRA FLATOW: And I’m very happy for that.
[LAUGHTER]
We all are. I want to thank my guests Dr. Debra Mathews, Assistant Director for Science Programs of the Berman Institute of Bioethics, Associate Professor of Pediatrics at Johns Hopkins. Dr. Jef Boeke, Director of the Institute of System Genetics, and Professor in Department of Biochemistry and Molecular Pharmacology at New York University Langone Health, right down the block here. Dr. Floyd Romesberg, Professor of Chemistry at the Scripps Research Institute in La Jolla, California. Thank you all for taking time to do this.
DEBRA MATHEWS: Thank you.
FLOYD ROMESBERG: My pleasure.
IRA FLATOW: One last thing before we go. Science Friday is hopping on the subway over to Brooklyn for an evening of science at BAM, Brooklyn Academy of Music. That’s on Saturday, April 27. We’re putting on a special live edition of Science Friday with music and demos, and we’re going to look at the secretly science lives of city pigeons. Don’t you wonder about how do city pigeons live and function? We’re going to tell you everything you ever wanted to know. And you’re not going to want to miss it. So you can join us Saturday, April 27 at BAM. Saturday– mark it in your calendar. It’s at night. Saturday night. April 27 at BAM. Tickets and info ScienceFriday.com/Brooklyn. See you there, as we say in Brooklyn.
Charles Bergquist is our director, and senior producer, Christopher Intagliata. Our producers are Alexa Lim, Christie Taylor, Katie Feather. Technical and engineering help today from Rich Kim, Sarah Fishman, and Kevin Wolfe. We’re active all week on Facebook, Twitter, Instagram, all social media. If you have a smart speaker, you can ask it to play Science Friday, if you can’t hear it on the radio whenever you want. So every day now is Science Friday. We’ll see you in Brooklyn, Saturday, April 27, at BAM. I’m Ira Flatow in New York.
Copyright © 2019 Science Friday Initiative. All rights reserved. Science Friday transcripts are produced on a tight deadline by 3Play Media. Fidelity to the original aired/published audio or video file might vary, and text might be updated or amended in the future. For the authoritative record of Science Friday’s programming, please visit the original aired/published recording. For terms of use and more information, visit our policies pages at http://www.sciencefriday.com/about/policies/
Alexa Lim was a senior producer for Science Friday. Her favorite stories involve space, sound, and strange animal discoveries.