The Genetic Roots Of Antibiotic Resistance
12:13 minutes
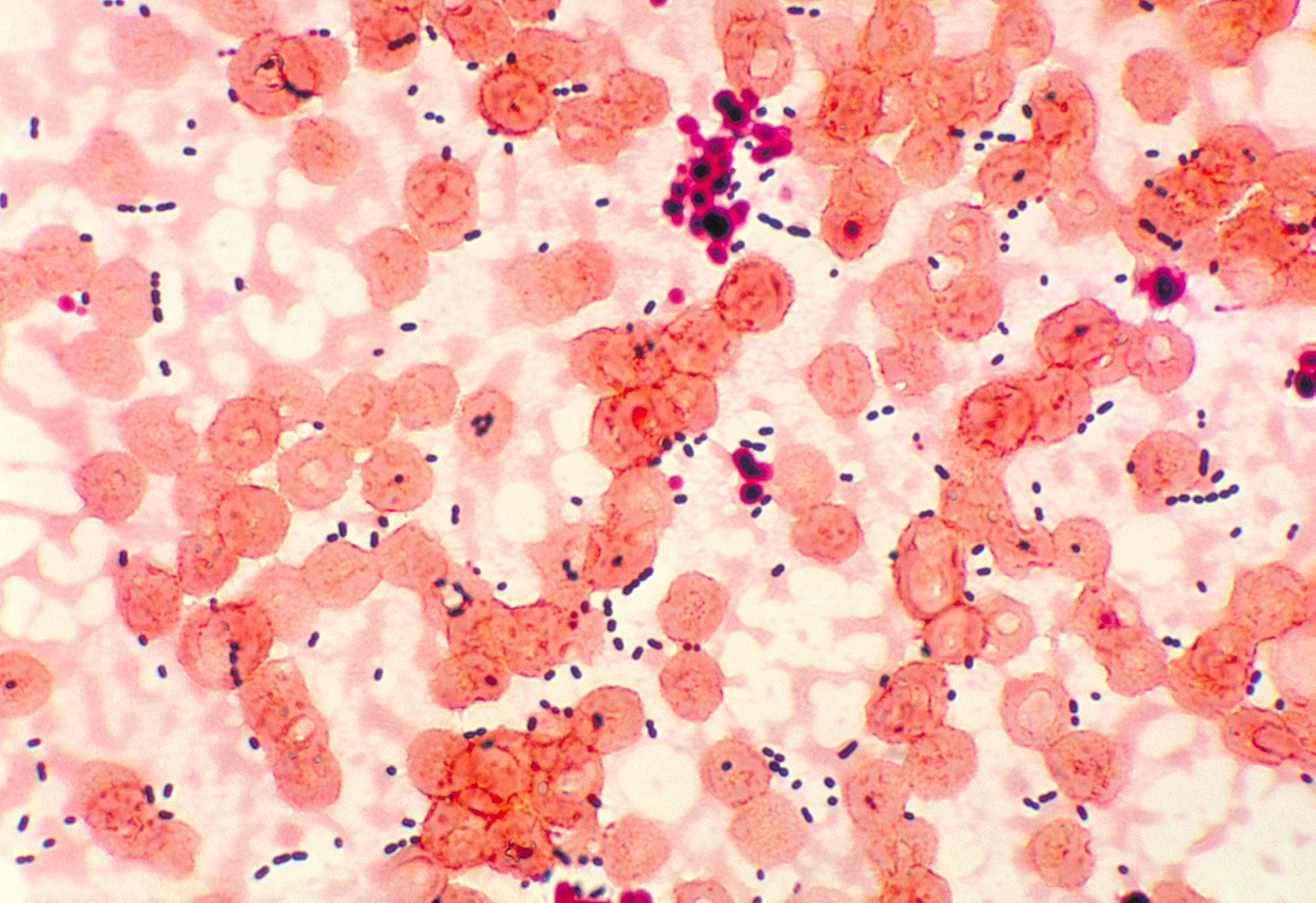
Antibiotic resistance—when pathogens no longer respond to the conventional antibiotic medications—is a serious medical problem. According to the CDC, over 2.8 million antibiotic-resistant infections occur in the U.S. each year, causing some 35,000 deaths. It’s in part due to overprescription of antibiotics in medicine, and the widespread use of antibiotics in animal agriculture. But the problem isn’t entirely of humans’ making. The roots of antibiotic resistance go back millions of years.
A recent study in the Proceedings of the National Academy of Sciences collected hundreds of soil and poop samples from around the world, to try to trace back the genetics of how resistance arose in Enterococcus, a genus of bacteria that live in the guts of pretty much every land animal. In the course of their analysis, the researchers identified 18 entirely new species in the genus Enterococcus, with over 1,000 genes that had never been seen before.
Dr. Michael Gilmore, the Chief Scientific Officer at Mass Eye and Ear, joins Ira to talk about the study and what the team hopes to learn about the causes of antibiotic resistance.
Invest in quality science journalism by making a donation to Science Friday.
Dr. Michael Gilmore is the Chief Scientific Officer of Mass Eye and Ear, and a professor at Harvard Medical School in Boston, Massachusetts.
IRA FLATOW: Antibiotic resistance is a serious medical problem. That’s when pathogens no longer respond to the conventional medications. It’s in part due to overprescription of antibiotics in medicine and the widespread use of antibiotics in animal agriculture.
But here’s something new. Turns out the roots of antibiotic resistance go way back before humans were around, millions of years, to the soil. A recent study in the proceedings of the National Academy of Sciences collected hundreds of soil and poop samples from around the world to try to trace back the genetics of how resistance arose in Enterococcus bacteria. That’s a group of bacteria that live in the guts of pretty much every land animal.
Joining me now is Dr. Michael Gilmore. He’s the chief scientific officer at Mass Eye and Ear and professor at Harvard Med School. Welcome to Science Friday.
MICHAEL GILMORE: Thanks so much, Ira. Pleasure to be here.
IRA FLATOW: Tell us about this bacteria. And why study it?
MICHAEL GILMORE: Yeah, so Enterococcus is especially interesting. In the time that we’ve used antibiotics on humans, antibiotic-resistant bacteria have emerged to cause problems. And two of those bacteria are members of the Enterococcus family. That’s out of thousands of possible bacteria.
So it caught our attention that Enterococcus arose, and it arose twice. So that said, there’s something special about this. So now it’s among the leading causes of antibiotic-resistant infection.
IRA FLATOW: So you’re trying to map out the genetics of many of these species to see how they fit together, right? How do you go about doing that?
MICHAEL GILMORE: Exactly. Genomics is the new lens to understand and view the world through. And that’s showing us so much that we didn’t understand before. When you look through the microscope at bacteria, they’ll look tiny and kind of round. And it’s hard to distinguish one from the other.
But when you look at the genomes, you can see every detail of the organism’s biology. We can’t completely understand it yet, but all the information is there. So we compare the genomes. And we ask, how far apart are they? It’s sort of like 23andMe on bacteria.
IRA FLATOW: So you collect them from around the world. Some of the more exotic ones come from where?
MICHAEL GILMORE: We teamed up with some elite adventurers, a group called Adventures and Scientists for Conservation. And their adventures go to places where humans have never been. And this included glaciers in Greenland, mountains in Nepal, a lot of different places.
IRA FLATOW: So they’re out there collecting poop samples that they send back to you in a mailer, so to speak?
MICHAEL GILMORE: [LAUGHS] Yes. We get all kinds of samples. We get dead insects. We get poop samples. Sometimes we get a piece of intestine or something like that. They’re appropriately labeled as biological specimens and put in the mail, but sometimes they get pretty stinky by the time they arrive.
IRA FLATOW: I can imagine there’s some grad students opening it, not you, right?
MICHAEL GILMORE: [LAUGHS] Yeah, that’s right. We need extreme adventures for that.
IRA FLATOW: [LAUGHS] I’m struck by the fact that these bacteria are in all sorts of animals, from people to penguins to insects to even some aquatic species, just how similar are all of them?
MICHAEL GILMORE: The human use of antibiotics on people and on animals has really changed the genomes of these bacteria. And that’s one of the things we learned from this study. When we look at the bacteria that come from human antibiotic-resistant infections, those Enterococci have genomes that are as much as 25% larger than the strains that are in the wild.
And that’s because they’ve collected all kinds of things in the effort to survive in the face of a lot of antibiotic use. Some of the things they collected are useful, some not as far as we can tell. But that’s one of the profound differences between hospital strains and strains from the wild.
Also, in the hospital, we mainly see two species. In the wild, our collection has shown that there are probably thousands of species. We just added 18 new species, bringing the total to about 80. But we would project, based on our rate of hit of new species, there are tens of thousands of species out there that we don’t know about.
IRA FLATOW: And what do you make use of? What does this tell you?
MICHAEL GILMORE: Well, a lot of things. So our interest is mainly trying to understand how antibiotic resistance emerged in Enterococcus. And Enterococcus got an early start when animals first crawled on land 425 or 450 million years ago. One of the things that distinguishes Enterococcus is they’re kind of like the cockroach of bacteria. They survive very harsh conditions.
So when the first invertebrates crawled onto land and pooped in the sand, a proto Enterococcus was in their gut. And it was probably one of the ones that survived the best on the sand and then re-entered another animal when it crawled on shore and was passed that way. So the ability to survive very harsh exposure is the hallmark of Enterococcus.
And turns out we work very, very hard to decontaminate hospital beds after patient use and things like that. But one of the challenges is Enterococcus. It’s super difficult to kill. So that, I think, is core to its biology, as well as the reason it emerged as a problem in hospitals.
IRA FLATOW: Now, in your research, you call out two species as being kind of unusual. And I’m talking about dragonflies and chickens. Wow. Why?
MICHAEL GILMORE: Yeah, so one of the ways we get Enterococci in our guts is from the food we eat. Chickens eat a lot of different things. They’re scratch feeders naturally. So they scratch the dirt and eat insects. And the guts of those insects have Enterococcus.
And those insects have lived in the soil where antibiotic producers naturally occur. And so we strongly suspect all of the antibiotic resistance as we see in hospital patients now originated in the soil ecosystem. And we think chickens, especially those fed antibiotics to promote their weight gain, collect Enterococci in their gut, and again, especially the antibiotic-resistant ones. And once they’re in chickens, then the chickens get processed and passed to humans. So we think it’s because of the variety of things chickens eat and that they come from the soil, where there’s tremendous diversity.
Dragonflies are also carnivores and for the same reason. They eat a lot of different types of insects. And we found a ton of Enterococcus diversity in insects.
IRA FLATOW: So we’re not just talking about antibiotics in the sense of a pill or a liquid you get at the drugstore. You’re talking about stuff that’s found naturally in nature.
MICHAEL GILMORE: Exactly. So antibiotics are about a billion years old. And the first organism that produced an antibiotic also had to be resistant to it. So resistances generally can be traced back to the original antibiotic producers.
But once that producer started producing it in the soil, when the producing organism died and spilled out its DNA, neighboring microbes collected some of that DNA and picked up some of the resistances. And the resistances got spread outward that way. So antibiotic production and antibiotic resistance is ancient. But it had always been confined to the soil ecosystem. It was only since the 1930 that it was brought into contact with humans in a substantial way.
IRA FLATOW: Have you found from your hundreds of soil and poop samples anything new there, anything that surprised you, new species or whatever?
MICHAEL GILMORE: Yes. So what surprised us is that we found 18 new species that had never been seen before. And that included over 1,000 new genes that also had never been seen before. And also in some species that we did know about, we found some new toxins. In collaboration with some of our associates here at Harvard Med School, we found a new type of botox. And we found another pore-forming toxin that has a very interesting novel mechanism.
IRA FLATOW: Huh. Do you need any fresh samples? I mean, should our listeners send you stuff?
MICHAEL GILMORE: We do. So now, Ira, this paper showed us where the unexplored diversity is. And it’s in insects and their invertebrate relatives. We really want to systematically explore that now. Whereas we’re up to about 80 species now, we think we can easily double that by examining maybe 5,000 or 10,000 invertebrate samples. And we now have the high-throughput genomics to do that.
IRA FLATOW: I go through a lot of natural history museums. And you see samples of all kinds of stuff they’re in jars or pinned to paper. Could they be sources that you could look into?
MICHAEL GILMORE: Yeah, that really is a great idea. And they certainly can be. The problem that we run into is we like to recover the live bacteria when we can so that when we find something interesting in its genome, we can culture the bacteria and explore its properties.
From preserved specimens, if they’re preserved in alcohol or something like that, the bacteria generally are dead. But we can work with the DNA. But we prefer just a dead insect. And again, since Enterococci are so persistent, the Enterococcus in a dead insect can survive in the mail for weeks.
IRA FLATOW: Whenever we talk about sampling and finding new stuff, I’m always struck by the amount of stuff that’s still out there. I mean, how much of what you have found do you think represents the total of what’s out there that we don’t know about?
MICHAEL GILMORE: That is the astounding thing. So our collection rate told us that we’re somewhere around 1%, knowing about 1% of what’s out there with respect to Enterococcus. And Enterococcus is one of the more common organisms. And so it boggles the mind, really, what’s left in that 99% that hasn’t been even ever seen before.
IRA FLATOW: Do you ever think about searching for phages, the viruses that attack bacteria?
MICHAEL GILMORE: Absolutely. That’s a very hot idea. So that’s something that we’ve done. Many of these do have phages. Some phages are sleeping in the genomes of many of these strains. And those phages can certainly be employed for the new kinds of therapies that are being explored right now.
IRA FLATOW: OK, so what do you want to look for next? What’s on your agenda?
MICHAEL GILMORE: Well, one of the things that we’d like to do is we know that antibiotic resistance genes originate in the soil. And we know that they end up in the hospital. We would like to put together the chain of custody in between those so that we know exactly what practices are promoting the upwelling of antibiotic resistance from the soil ecosystem so that we can effectively stop it.
We don’t want to just have a blanket moratorium on use of antibiotics. Then the companies go out of business. We want to strategically deploy antibiotics in a way that doesn’t promote bringing in new resistances from the soil ecosystem. And so we need to know how it moves. And so that’s why we’re out exploring nature and trying to connect the dots.
IRA FLATOW: Well, Dr. Gilmore, thank you for connecting the dots for us.
MICHAEL GILMORE: A real pleasure, Ira. I enjoyed it, and I’m a big fan.
IRA FLATOW: Thank you, Dr. Michael Gilmore. He’s the chief scientific officer at Mass Eye and Ear and a professor at Harvard Med School.
Copyright © 2023 Science Friday Initiative. All rights reserved. Science Friday transcripts are produced on a tight deadline by 3Play Media. Fidelity to the original aired/published audio or video file might vary, and text might be updated or amended in the future. For the authoritative record of Science Friday’s programming, please visit the original aired/published recording. For terms of use and more information, visit our policies pages at http://www.sciencefriday.com/about/policies/
As Science Friday’s director and senior producer, Charles Bergquist channels the chaos of a live production studio into something sounding like a radio program. Favorite topics include planetary sciences, chemistry, materials, and shiny things with blinking lights.
Ira Flatow is the founder and host of Science Friday. His green thumb has revived many an office plant at death’s door.